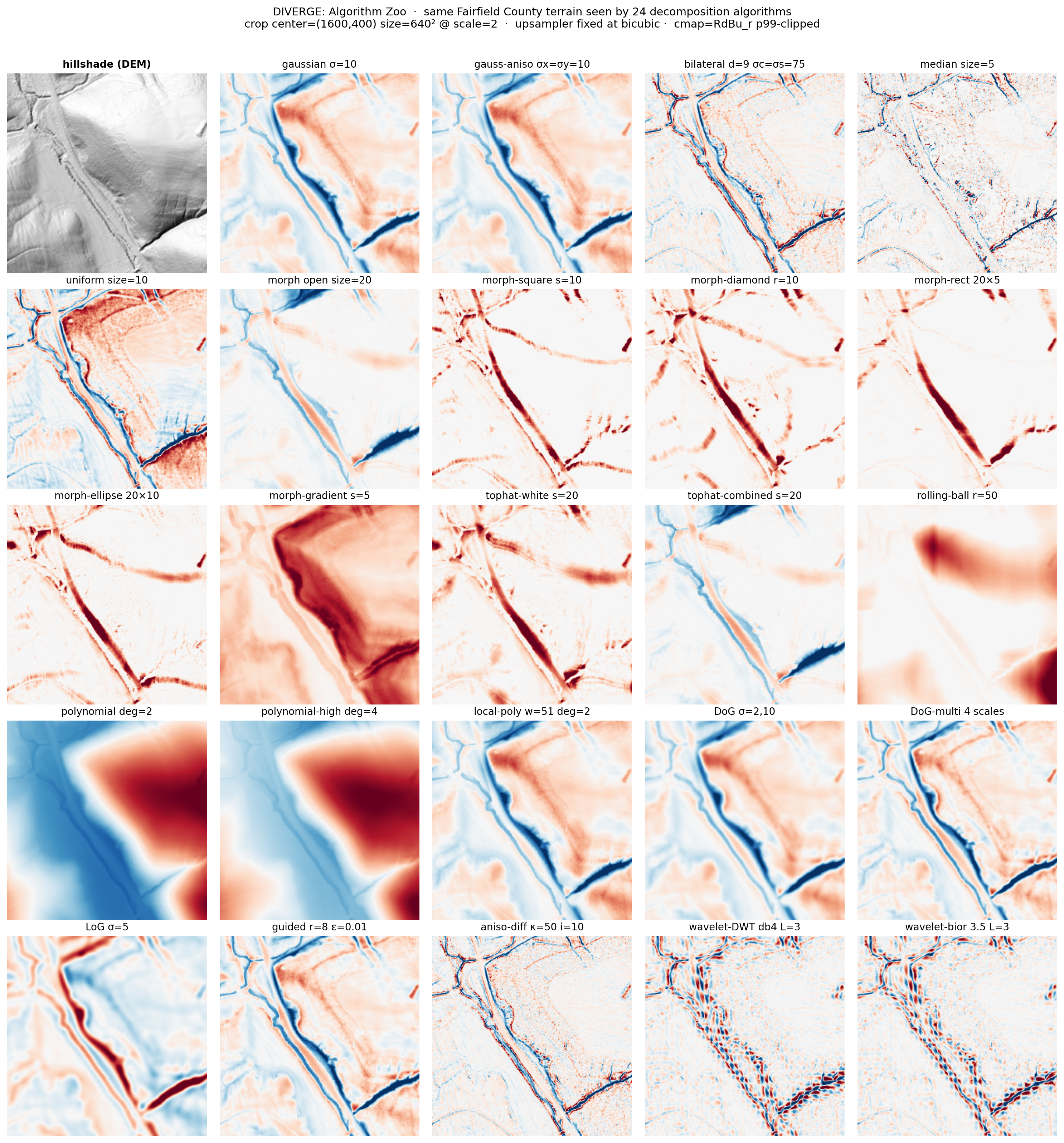
The algorithm zoo
The same 640×640 patch of Fairfield County terrain — a sweeping diagonal stream channel with branching drainage and a sharp V-shaped linear feature — seen through 24 different decomposition algorithms, all upsampled with bicubic_scale2 for fair comparison. Each one preserves and destroys different feature types. Wavelets become watered silk. Morphological-rect carves out V-shapes. Polynomial trend-removal paints regional gradients. Bilateral becomes a comic-book inkwash.